High-resolution immune repertoire insights
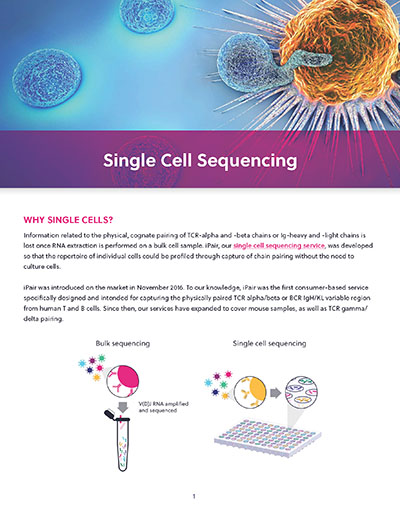
Single cell sequencing
for CAR-T cell therapy development & immunotherapy trials
A patient’s unique profile of B and T cells determines their ability to neutralize cancer cells and is predictive of their response to immunotherapies. But typical immune cell sequencing approaches do not preserve crucial information related to the cognate pairing of receptor chains and fail to capture critically important rare cell types.
![]()
Single cell sequencing and transcriptome profiling can be used to screen cancer patient samples for specific cellular biomarkers, such as PD1 and CTLA-4, to identify patients likely to respond in clinical trials. Single cell analysis can also be used to assess CAR-T cell therapy development, track cellular clonality and evaluate immune cell activation following antigen stimulation. High-resolution profiling may provide clues into patterns of variable immune cell activity and immunotherapy response.

Questions about iPair single cell sequencing?
Case Study
Cell therapy response
Learn how single cell TCR sequencing supports
- Identify paired receptor sequences for downstream CAR-T development or validation purposes
- Characterize progression-free survival following neoantigen treatment
- Track expansion of tumor specific cytotoxic T cells
- Explore the role of TCR diversity in improving clinical outcomes

Ready to get started?
iPair/iPair + services
iRepertoire serves as a leader in the field of immune repertoire sequencing. We invite you to consult with our team of scientists to discover how iPair/iPair+ services can accelerate your analysis of immune cell function and diversity.
Schedule a consultation for single cell sequencing.