Learning Center
Why sequence the immune repertoire?
“Our immune system is the smartest and best doctor around, and if we can learn from the best, we can be better.” Dr. Jian Han, founder and CSO of iRepertoire Sequencing the expressed T and B cell receptor genes in the immune repertoire provides a detailed picture of a person’s health. The amount of diversity…
Read MoreHow to perform immune repertoire sequencing
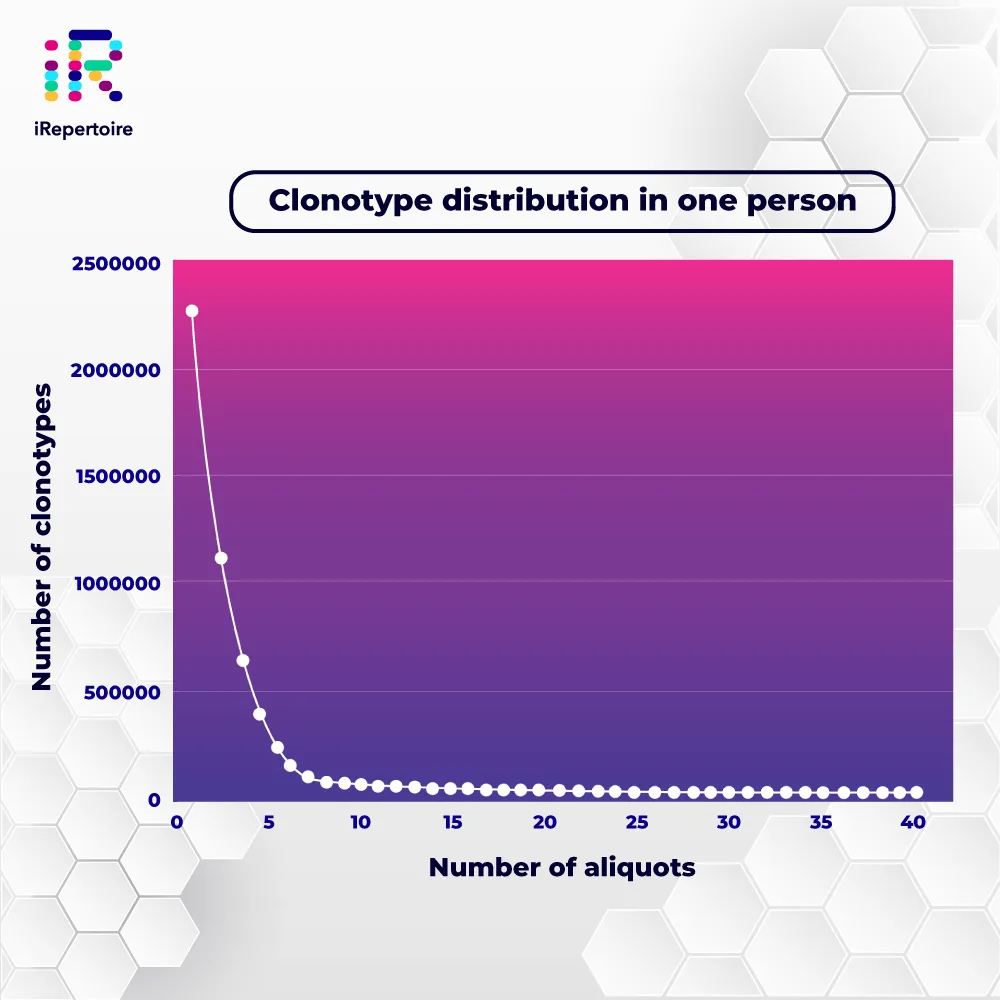
Capturing the full clonal diversity of the immune repertoire can reveal important information about the pathology and prognosis of disease. Learn how now.
Read MoreiRepertoire’s primer systems
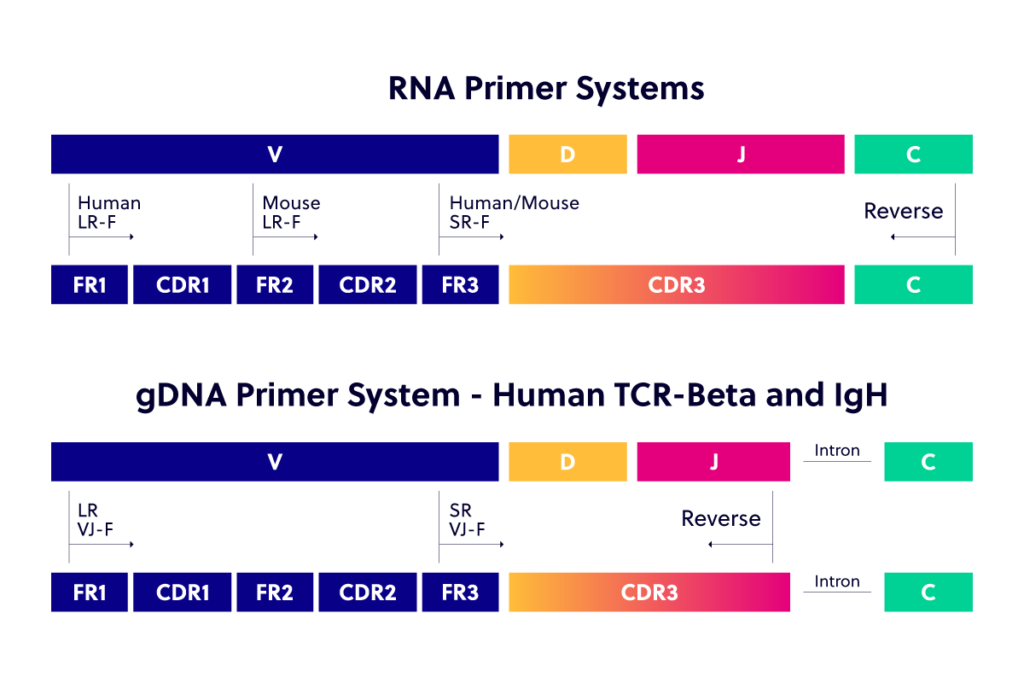
iRepertoire has developed multiplex PCR primer mixes for V(D)J amplification and sequencing for both our arm-PCR and dam-PCR technologies. In these multiplex PCR systems, a nested set of inside and outside primers has been designed and optimized for every V(D)J target needing to be co-amplified in the multiplex assay. We offer primers, amplification services, and…
Read MoreiRepertoire’s amplification technologies
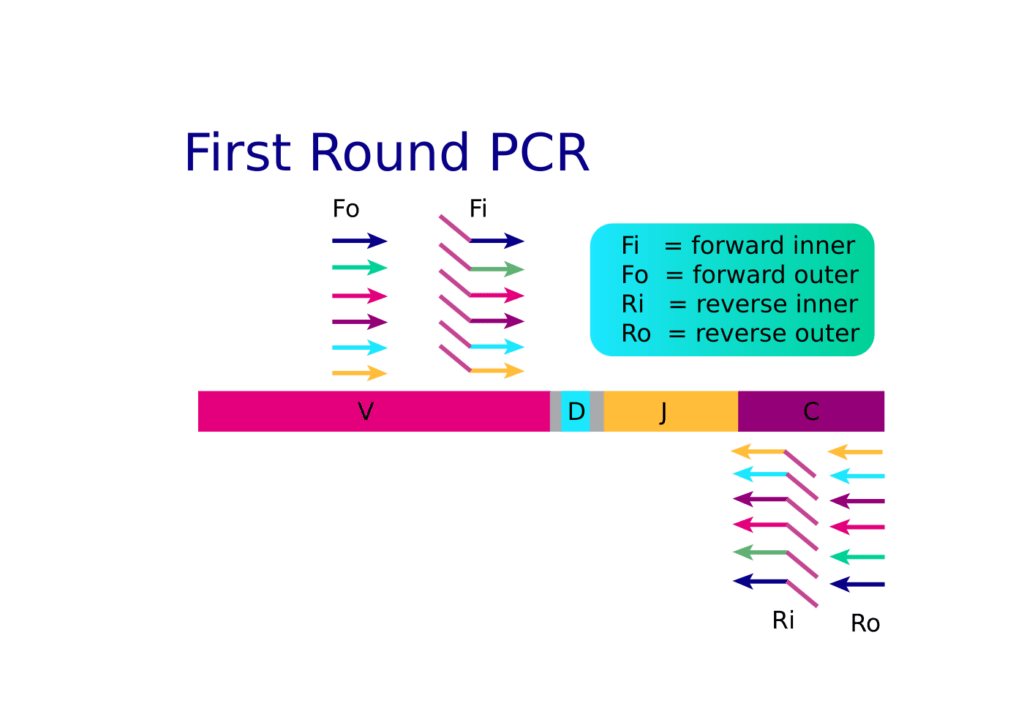
Challenges of immune repertoire sequencing The unique challenges involved with sequencing the immune repertoire have necessitated the development of repertoire-specific library preparation technologies. In order to fully capture the diversity of the immune repertoire, amplification technologies have to be used that account for biases towards small amplicons and amplifying rare clonotypes beyond the templates that…
Read MoreiRepertoire’s immune sequencing services
Immune repertoire sequencing: RepSeq and RepSeq+ iRepertoire’s sequencing services RepSeq and RepSeq+ complement our amplification technologies arm-PCR and dam-PCR, respectively. Because arm-PCR and dam-PCR have different strengths, the decision of whether to use RepSeq or RepSeq+ will depend on the nature of your specific project. Arm-PCR uses internal and external nested primers for extremely inclusive…
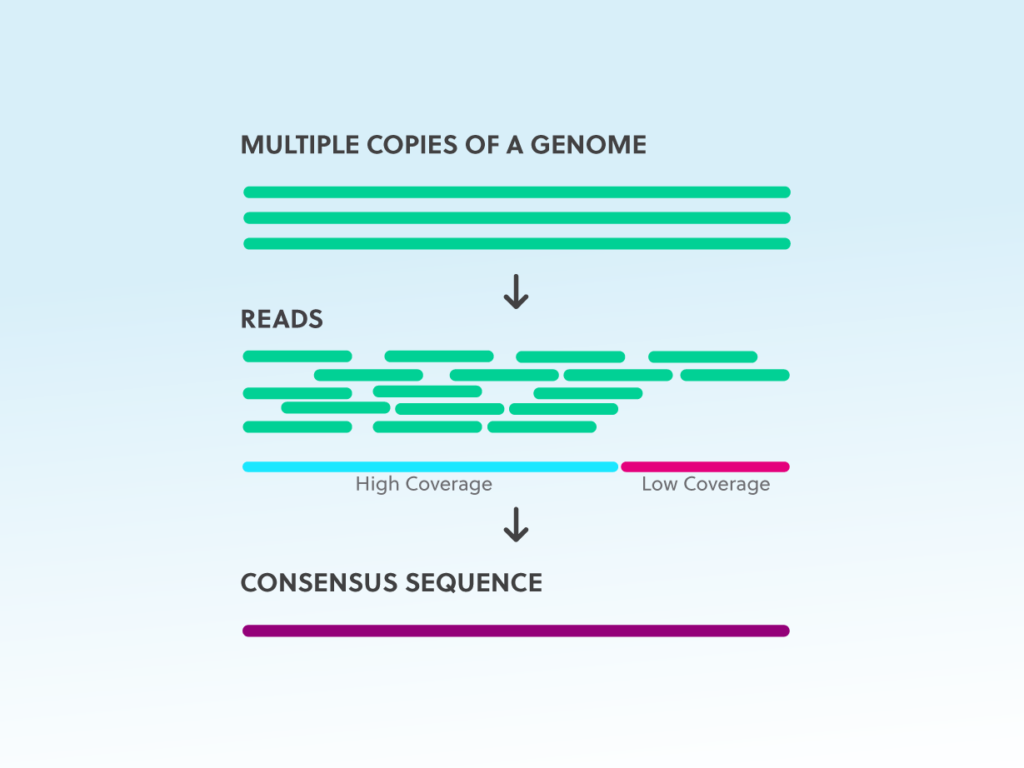
Read MoreNGS considerations: coverage, read length, multiplexing
Plan your NGS experiment to adjust cost and amount of sequencing data to best answer your experimental questions. Dive into the top NGS considerations here.
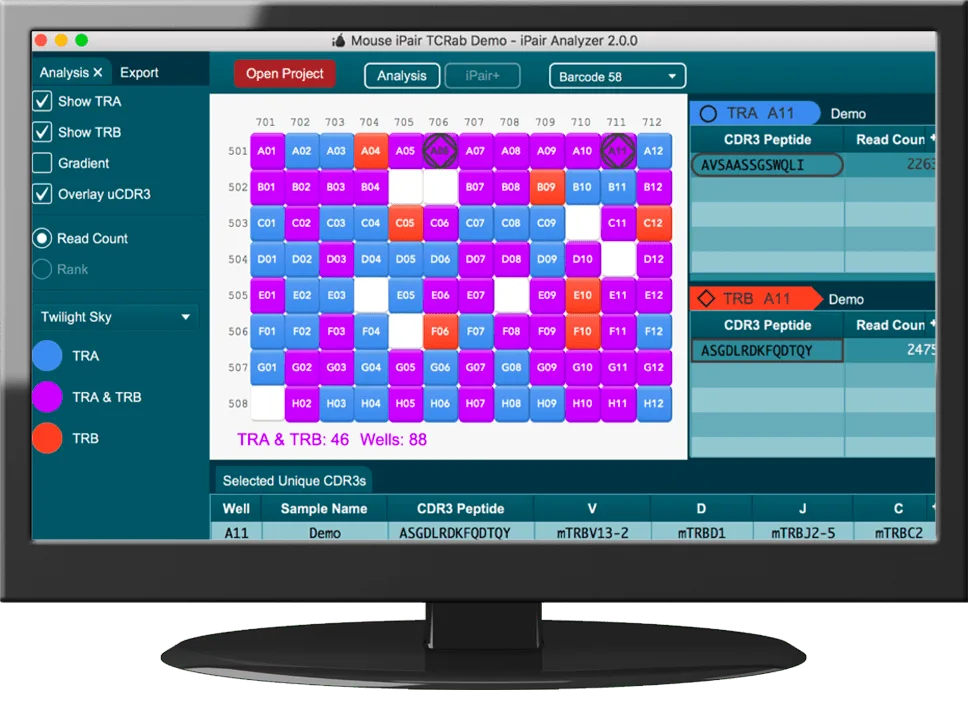
Read MoreiPair: iRepertoire’s single cell immune sequencing service
Why single cells? Information related to the physical, cognate pairing of TCR-alpha and -beta chains or Ig-heavy and -light chains is lost once RNA extraction is performed on a bulk cell sample. iPair, our single cell immune sequencing service, was developed so that the repertoire of individual cells could be profiled through capture of chain…
Read MoreiRepertoire’s data analysis services
All of our services come with complimentary basic data analysis through our proprietary pipeline, which then outputs data to iRweb. iRweb is a web-based RepSeq bioinformatic software that allows researchers to have all of the following services and analyses performed on their samples: Barcode demultiplexing The number of CDR3s captured in the library and the…
Read MoreiRweb: Technical notes
Overview This page describes the technical steps involved in iRweb’s bioinformatic pipeline. If you are unfamiliar with data analysis for next-generation sequencing, we suggest you visit our NGS overview page. For a general introduction to iRweb, see our Learning Center article about data analysis. For more details about iRweb than what can be found on…
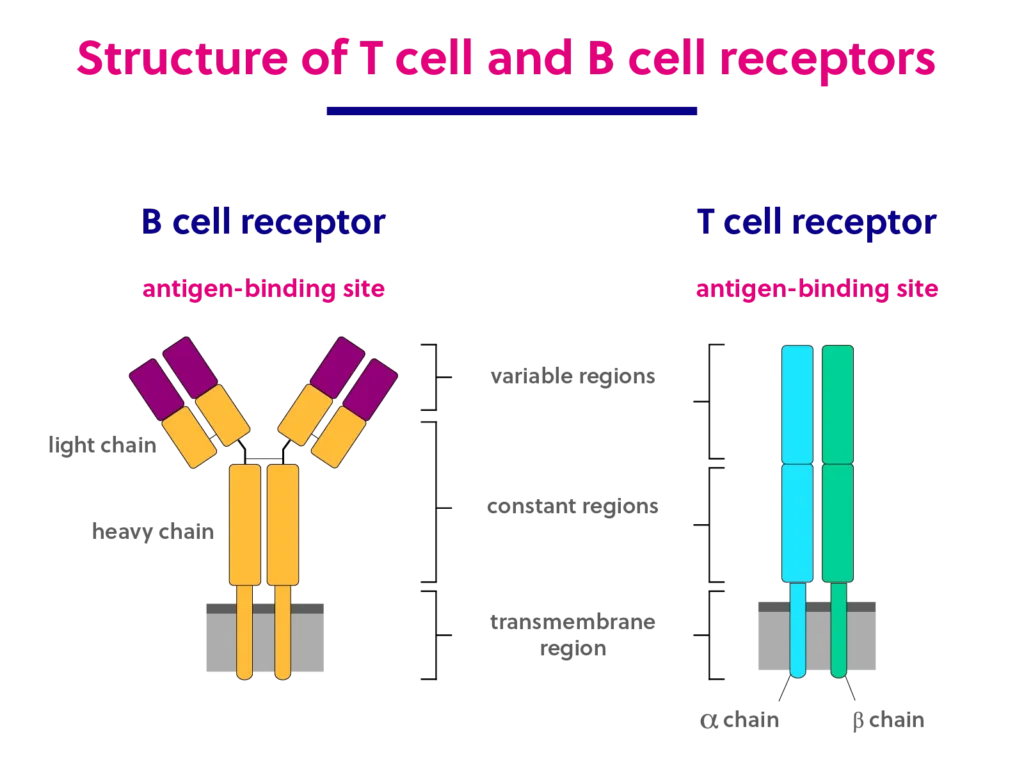
Read MoreB cell and T cell structure and function
Introduction to T cells and B cells T cell and B cell lymphocytes work together to recognize foreign substances called antigens. As the primary agents responsible for adaptive immunity, T cells and B cells are sometimes called the “special ops” of the immune system. Inherent structural features of the B and T cell receptors are…
Read More